10-11易位蛋白2(TET2)介导的表观遗传与肠道微生物-免疫互作对自身免疫性肝炎的调控作用
DOI: 10.12449/JCH260327
Regulatory effect of ten-eleven translocation 2-mediated epigenetics and the interaction between gut microbiota and immunity on autoimmune hepatitis
-
摘要: 10-11易位蛋白2(TET2)作为表观遗传调控核心酶,可通过介导DNA去甲基化动态调节CD4+ T细胞分化及功能。近年研究发现,TET2缺失可通过扰乱辅助性T细胞17/调节性T细胞之间的平衡,激活肠-肝轴炎症信号通路,从而促进自身免疫性肝炎(AIH)进展。本文系统综述了TET2在CD4+ T细胞和肠道微生物间的桥梁作用,深入探讨其通过肠道微生物-表观遗传-免疫网络驱动AIH的分子机制,并展望靶向TET2-菌群轴的潜在干预策略。Abstract: Ten-eleven translocation 2 (TET2), as a core enzyme in epigenetic regulation, dynamically regulates the differentiation and function of CD4+ T cells by mediating DNA demethylation. Recent studies have shown that TET2 deficiency can promote the progression of autoimmune hepatitis (AIH) by disrupting the Th17/Treg balance and activating inflammatory signals along the gut-liver axis. This article systematically reviews the bridging role of TET2 between CD4+ T cells and gut microbiota, explores the molecular mechanisms by which it drives AIH through the gut microbiota-epigenetics-immunity network, and discusses the potential intervention strategies targeting the TET2-microbiota axis.
-
Key words:
- Hepatitis, Autoimmune /
- Epigenomics /
- Gastrointestinal Microbiome
-
注: TET2,10-11易位蛋白2;TLR4,Toll样受体4;MyD88,髓系分化初级反应蛋白质88;NF-κB, 核因子κB;miRNA,微RNA;5mC,5-甲基胞嘧啶;5hmC,5-羟甲基胞嘧啶;oxi-mC,氧化修饰胞嘧啶;5fC,5-甲酰基胞嘧啶;5caC,5-羧基胞嘧啶;TGF-β,转化生长因子β;IL,白细胞介素;Treg,调节性T细胞;Th17,辅助性T细胞17;HC,肝细胞;HSC,肝星状细胞;FOXP3,叉头框蛋白p3;STAT3,信号转导与转录激活因子3;RoRγt,视黄酸受体相关孤儿受体γt;GPR43,G蛋白偶联受体43;cAMP,环磷酸腺苷;PKA,环磷酸腺苷依赖性蛋白激酶A;AIH,自身免疫性肝炎。
图 1 肠道微生物-TET2-CD4+T细胞轴驱动AIH发展的分子机制
Figure 1. Molecular mechanism of the gut microbiota-TET2-CD4+T cell axis driving the development of AIH
-
[1] HARTL J, MIQUEL R, ZACHOU K, et al. Features and outcome of AIH patients without elevation of IgG[J]. JHEP Rep, 2020, 2( 3): 100094. DOI: 10.1016/j.jhepr.2020.100094. [2] STOELINGA AEC, BIEWENGA M, DRENTH JPH, et al. Diagnostic criteria and long-term outcomes in AIH-PBC variant syndrome under combination therapy[J]. JHEP Rep, 2024, 6( 7): 101088. DOI: 10.1016/j.jhepr.2024.101088. [3] LAPIERRE P, ALVAREZ F. Type 2 autoimmune hepatitis: Genetic susceptibility[J]. Front Immunol, 2022, 13: 1025343. DOI: 10.3389/fimmu.2022.1025343. [4] WANG L, CAO ZM, ZHANG LL, et al. The role of gut microbiota in some liver diseases: From an immunological perspective[J]. Front Immunol, 2022, 13: 923599. DOI: 10.3389/fimmu.2022.923599. [5] CHEN HR, HAN ZY, FAN YY, et al. CD4+ T-cell subsets in autoimmune hepatitis: A review[J]. Hepatol Commun, 2023, 7( 10): e0269. DOI: 10.1097/HC9.0000000000000269. [6] SHEN MY, ZHOU LY, FAN XL, et al. Metabolic reprogramming of CD4+ T cells by mesenchymal stem cell-derived extracellular vesicles attenuates autoimmune hepatitis through mitochondrial protein transfer[J]. Int J Nanomedicine, 2024, 19: 9799- 9819. DOI: 10.2147/IJN.S472086. [7] VUERICH M, WANG N, KALBASI A, et al. Dysfunctional immune regulation in autoimmune hepatitis: From pathogenesis to novel therapies[J]. Front Immunol, 2021, 12: 746436. DOI: 10.3389/fimmu.2021.746436. [8] MUSCATE F, WOESTEMEIER A, GAGLIANI N. Functional heterogeneity of CD4+ T cells in liver inflammation[J]. Semin Immunopathol, 2021, 43( 4): 549- 561. DOI: 10.1007/s00281-021-00881-w. [9] SIRBE C, SIMU G, SZABO I, et al. Pathogenesis of autoimmune hepatitis-cellular and molecular mechanisms[J]. Int J Mol Sci, 2021, 22( 24): 13578. DOI: 10.3390/ijms222413578. [10] ZHANG KH, LI JC, SHI Z, et al. Ginsenosides regulates innate immunity to affect immune microenvironment of AIH through hippo-YAP/TAZ signaling pathway[J]. Front Immunol, 2022, 13: 851560. DOI: 10.3389/fimmu.2022.851560. [11] LI YY. Modern epigenetics methods in biological research[J]. Methods, 2021, 187: 104- 113. DOI: 10.1016/j.ymeth.2020.06.022. [12] ÄIJÖ T, THEOFILATOS D, CHENG M, et al. TET proteins regulate T cell and iNKT cell lineage specification in a TET2 catalytic dependent manner[J]. Front Immunol, 2022, 13: 940995. DOI: 10.3389/fimmu.2022.940995. [13] GIOULBASANI M, ÄIJÖ T, LIU SY, et al. Concomitant loss of TET2 and TET3 results in T cell expansion and genomic instability in mice[J]. Commun Biol, 2024, 7( 1): 1606. DOI: 10.1038/s42003-024-07312-0. [14] CONG BY, ZHANG Q, CAO XT. The function and regulation of TET2 in innate immunity and inflammation[J]. Protein Cell, 2021, 12( 3): 165- 173. DOI: 10.1007/s13238-020-00796-6. [15] ZACHOU K, ARVANITI P, LYBEROPOULOU A, et al. Altered DNA methylation pattern characterizes the peripheral immune cells of patients with autoimmune hepatitis[J]. Liver Int, 2022, 42( 6): 1355- 1368. DOI: 10.1111/liv.15176. [16] BAESSLER A, NOVIS CL, SHEN ZL, et al. Tet2 coordinates with Foxo1 and Runx1 to balance T follicular helper cell and T helper 1 cell differentiation[J]. Sci Adv, 2022, 8( 24): eabm4982. DOI: 10.1126/sciadv.abm4982. [17] BAESSLER A, FUCHS B, PERKINS B, et al. Tet2 deletion in CD4+ T cells disrupts Th1 lineage commitment in memory cells and enhances T follicular helper cell recall responses to viral rechallenge[J]. Proc Natl Acad Sci USA, 2023, 120( 36): e2218324120. DOI: 10.1073/pnas.2218-324120. [18] CHEN J, LIU W, ZHU WJ. FOXP3+ Treg cells are associated with pathological process of autoimmune hepatitis by activating methylation modification in autoimmune hepatitis patients[J]. Med Sci Monit, 2019, 25: 6204- 6212. DOI: 10.12659/MSM.915408. [19] PRETI M, SCHLOTT L, LÜBBERING D, et al. Failure of thymic deletion and instability of autoreactive Tregs drive autoimmunity in immune-privileged liver[J]. JCI Insight, 2021, 6( 6): e141462. DOI: 10.1172/jci.insight.141462. [20] OHKURA N, SAKAGUCHI S. Transcriptional and epigenetic basis of Treg cell development and function: Its genetic anomalies or variations in autoimmune diseases[J]. Cell Res, 2020, 30( 6): 465- 474. DOI: 10.1038/s41422-020-0324-7. [21] BAI L, HAO XL, KEITH J, et al. DNA methylation in regulatory T cell differentiation and function: Challenges and opportunities[J]. Biomolecules, 2022, 12( 9): 1282. DOI: 10.3390/biom12091282. [22] YUE XJ, LIO CJ, SAMANIEGO-CASTRUITA D, et al. Loss of TET2 and TET3 in regulatory T cells unleashes effector function[J]. Nat Commun, 2019, 10( 1): 2011. DOI: 10.1038/s41467-019-09541-y. [23] PACINI G, PAOLINO S, ANDREOLI L, et al. Epigenetics, pregnancy and autoimmune rheumatic diseases[J]. Autoimmun Rev, 2020, 19( 12): 102685. DOI: 10.1016/j.autrev.2020.102685. [24] WANG QX, SELMI C, ZHOU XM, et al. Epigenetic considerations and the clinical reevaluation of the overlap syndrome between primary biliary cirrhosis and autoimmune hepatitis[J]. J Autoimmun, 2013, 41: 140- 145. DOI: 10.1016/j.jaut.2012.10.004. [25] MIGITA K, KOMORI A, KOZURU H, et al. Circulating microRNA profiles in patients with type-1 autoimmune hepatitis[J]. PLoS One, 2015, 10( 11): e 0136908.10.1371/journal.pone. 0136908. [26] PANDEY SP, BENDER MJ, MCPHERSON AC, et al. Tet2 deficiency drives liver microbiome dysbiosis triggering Tc1 cell autoimmune hepatitis[J]. Cell Host Microbe, 2022, 30( 7): 1003- 1019. DOI: 10.1016/j.chom.2022.05.006. [27] ZHANG GL, WU SS, XIA GT. miR-326 sponges TET2 triggering imbalance of Th17/Treg differentiation to exacerbate pyroptosis of hepatocytes in concanavalin A-induced autoimmune hepatitis[J]. Ann Hepatol, 2024, 29( 2): 101183. DOI: 10.1016/j.aohep.2023.101183. [28] TORP AUSTVOLL C, GALLO V, MONTAG D. Health impact of the Anthropocene: The complex relationship between gut microbiota, epigenetics, and human health, using obesity as an example[J]. Glob Health Epidemiol Genom, 2020, 5: e2. DOI: 10.1017/gheg.2020.2. [29] SUN HJ, GUO YK, WANG HD, et al. Gut commensal Parabacteroides distasonis alleviates inflammatory arthritis[J]. Gut, 2023, 72( 9): 1664- 1677. DOI: 10.1136/gutjnl-2022-327756. [30] LIU JZ, GUO M, YUAN XB, et al. Gut microbiota and their metabolites: The hidden driver of diabetic nephropathy? Unveiling gut microbe’s role in DN[J]. J Diabetes, 2025, 17( 4): e70068. DOI: 10.1111/1753-0407.70068. [31] TZENG HT, LEE WC. Impact of transgenerational nutrition on nonalcoholic fatty liver disease development: Interplay between gut microbiota, epigenetics and immunity[J]. Nutrients, 2024, 16( 9): 1388. DOI: 10.3390/nu16091388. [32] SZABO G, BALA S, PETRASEK J, et al. Gut-liver axis and sensing microbes[J]. Dig Dis, 2010, 28( 6): 737- 744. DOI: 10.1159/000324281. [33] WANG H, WANG GD, BANERJEE N, et al. Aberrant gut microbiome contributes to intestinal oxidative stress, barrier dysfunction, inflammation and systemic autoimmune responses in MRL/lpr mice[J]. Front Immunol, 2021, 12: 651191. DOI: 10.3389/fimmu.2021.651191. [34] CHEN Y, GAN YM, ZHONG HJ, et al. Gut microbe and hepatic macrophage polarization in non-alcoholic fatty liver disease[J]. Front Microbiol, 2023, 14: 1285473. DOI: 10.3389/fmicb.2023.1285473. [35] ZHANG HX, LIU M, LIU X, et al. Bifidobacterium animalis ssp. lactis 420 mitigates autoimmune hepatitis through regulating intestinal barrier and liver immune cells[J]. Front Immunol, 2020, 11: 569104. DOI: 10.3389/fimmu.2020.569104. [36] KANG YB, KUANG XY, YAN H, et al. A novel synbiotic alleviates autoimmune hepatitis by modulating the gut microbiota-liver axis and inhibiting the hepatic TLR4/NF-κB/NLRP3 signaling pathway[J]. mSystems, 2023, 8( 2): e01127-22. DOI: 10.1128/msystems.01127-22. [37] WOO V, ALENGHAT T. Epigenetic regulation by gut microbiota[J]. Gut Microbes, 2022, 14( 1): 2022407. DOI: 10.1080/19490976.2021.2022407. [38] LV JL, HAO PD, ZHOU Y, et al. Role of the intestinal flora-immunity axis in the pathogenesis of rheumatoid arthritis-mechanisms regulating short-chain fatty acids and Th17/Treg homeostasis[J]. Mol Biol Rep, 2025, 52( 1): 617. DOI: 10.1007/s11033-025-10714-w. [39] LYON P, STRIPPOLI V, FANG B, et al. B vitamins and one-carbon metabolism: Implications in human health and disease[J]. Nutrients, 2020, 12( 9): 2867. DOI: 10.3390/nu12092867. [40] ANSARI I, RADDATZ G, GUTEKUNST J, et al. The microbiota programs DNA methylation to control intestinal homeostasis and inflammation[J]. Nat Microbiol, 2020, 5( 4): 610- 619. DOI: 10.1038/s41564-019-0659-3. [41] FERENC K, SOKAL-DEMBOWSKA A, HELMA K, et al. Modulation of the gut microbiota by nutrition and its relationship to epigenetics[J]. Int J Mol Sci, 2024, 25( 2): 1228. DOI: 10.3390/ijms25021228. [42] AMES SR, LOTOSKI LC, AZAD MB. Comparing early life nutritional sources and human milk feeding practices: Personalized and dynamic nutrition supports infant gut microbiome development and immune system maturation[J]. Gut Microbes, 2023, 15( 1): 2190305. DOI: 10.1080/19490976.2023.2190305. [43] ALIPOUR B, HOMAYOUNI-RAD A, VAGHEF-MEHRABANY E, et al. Effects of Lactobacillus casei supplementation on disease activity and inflammatory cytokines in rheumatoid arthritis patients: A randomized double-blind clinical trial[J]. Int J Rheum Dis, 2014, 17( 5): 519- 527. DOI: 10.1111/1756-185X.12333. [44] MA ZY, AKHTAR M, PAN H, et al. Fecal microbiota transplantation improves chicken growth performance by balancing jejunal Th17/Treg cells[J]. Microbiome, 2023, 11( 1): 137. DOI: 10.1186/s40168-023-01569-z. [45] CHEN L, RUAN GC, CHENG Y, et al. The role of Th17 cells in inflammatory bowel disease and the research progress[J]. Front Immunol, 2022, 13: 1055914. DOI: 10.3389/fimmu.2022.1055914. [46] WU Q, GE Z, LV C, et al. Interacting roles of gut microbiota and T cells in the development of autoimmune hepatitis[J]. Front Immunol, 2025, 16: 1584001. DOI: 10.3389/fimmu.2025.1584001. [47] MA HL, GAO WS, SUN XX, et al. STAT5 and TET2 cooperate to regulate FOXP3-TSDR demethylation in CD4+ T cells of patients with colorectal cancer[J]. J Immunol Res, 2018, 2018: 6985031. DOI: 10.1155/2018/6985031. [48] GHORESCHI K, LAURENCE A, YANG XP, et al. T helper 17 cell heterogeneity and pathogenicity in autoimmune disease[J]. Trends Immunol, 2011, 32( 9): 395- 401. DOI: 10.1016/j.it.2011.06.007. [49] HU X, ZOU YL, COPLAND DA, et al. Epigenetic drug screen identified IOX1 as an inhibitor of Th17-mediated inflammation through targeting TET2[J]. EBioMedicine, 2022, 86: 104333. DOI: 10.1016/j.ebiom.2022.104333. -



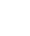 PDF下载 ( 1161 KB)
PDF下载 ( 1161 KB)
 下载:
下载:


