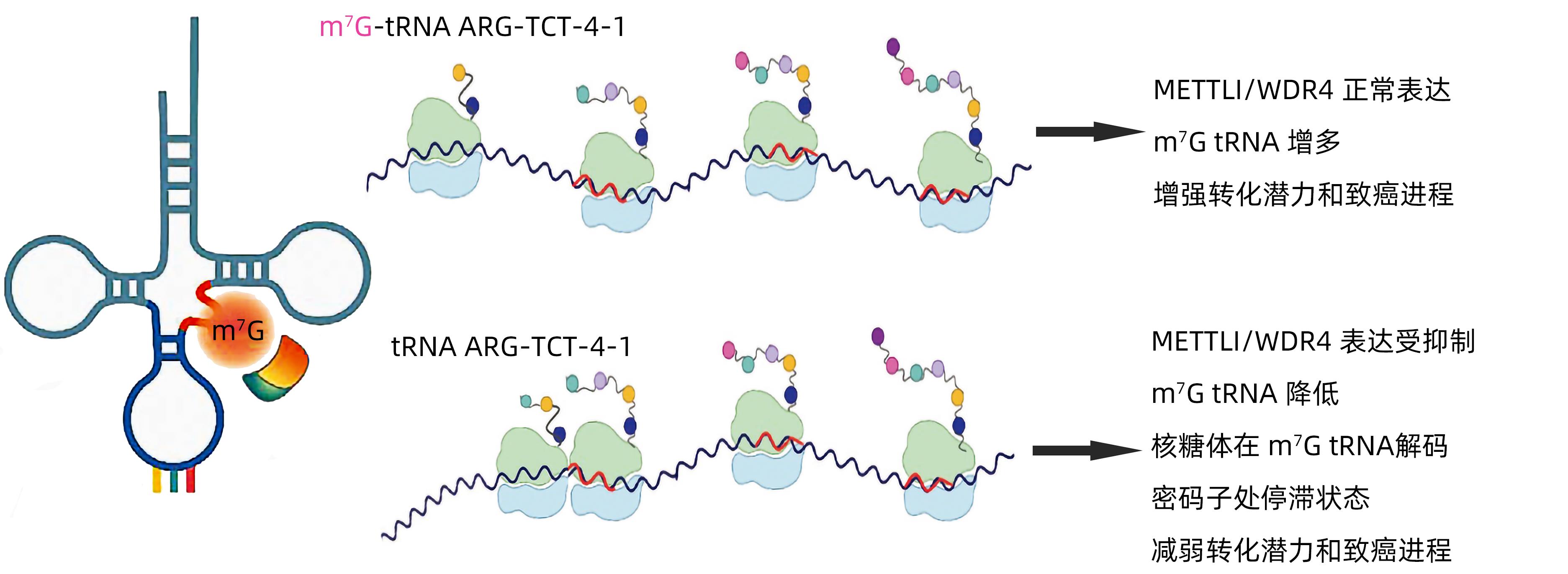
N7-甲基鸟苷(m7G)在肝细胞癌中的作用机制
DOI: 10.3969/j.issn.1001-5256.2023.12.029
利益冲突声明:本文不存在任何利益冲突。
作者贡献声明:高春负责论文撰写与修改;江晶晶、陈玉春等负责论文修改和审阅;郑晓凤负责数据收集;张久聪、于晓辉等负责拟定写作思路,指导撰写文章并最后定稿。
-
摘要: N7-甲基鸟苷(m7G)是目前最流行的RNA修饰之一,最近引起了国内外学者极大的关注。m7G修饰通过影响各种RNA分子(包括信使RNA、核糖体RNA、微RNA和转移RNA)的代谢,积极参与细胞的增殖、分化和凋亡等多种生物学过程。越来越多的证据表明,m7G尤其在癌症中起着关键作用,异常的m7G水平通过调节多个癌基因和抑癌基因的表达与肿瘤的发生和发展密切相关。肝细胞癌是我国最常见的消化道肿瘤,治疗效果欠佳。目前,肝细胞癌中m7G修饰的潜在分子机制尚不完全清楚。本文就m7G修饰在肝细胞癌中的潜在作用机制,以及m7G相关的诊断和治疗策略进行综述。Abstract: N7-methylguanosine (m7G) is one of the most popular RNA modifications at present and has attracted wide attention from researchers in China and globally. By influencing the metabolism of various RNA molecules (including messenger RNA, ribosomal RNA, microRNA, and transfer RNA), m7G modification actively participates in many biological processes such as cell proliferation, differentiation, and apoptosis. More and more evidence has shown that m7G plays a key role in the development of cancer, and abnormal m7G levels are closely associated with the development and progression of cancer by regulating the expression of multiple oncogenes and tumor suppressor genes. Hepatocellular carcinoma is the most common gastrointestinal tumor in China, and current treatment methods tend to have an unsatisfactory therapeutic effect. At present, the potential molecular mechanism of m7G modification in hepatocellular carcinoma remains unclear. This article reviews the potential mechanism of action of m7G modification in hepatocellular carcinoma and the m7G-related diagnosis and treatment strategies.
-
Key words:
- Carcinoma, Hepatocellular /
- N7-methylguanosine /
- Methylation
-
-
[1] SUNG H, FERLAY J, SIEGEL RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71( 3): 209- 249. DOI: 10.3322/caac.21660. [2] GANESAN P, KULIK LM. Hepatocellular carcinoma: New developments[J]. Clin Liver Dis, 2023, 27( 1): 85- 102. DOI: 10.1016/j.cld.2022.08.004. [3] REIG M, FORNER A, RIMOLA J, et al. BCLC strategy for prognosis prediction and treatment recommendation: The 2022 update[J]. J Hepatol, 2022, 76( 3): 681- 693. DOI: 10.1016/j.jhep.2021.11.018. [4] THAPAR R, BACOLLA A, OYENIRAN C, et al. RNA modifications: Reversal mechanisms and cancer[J]. Biochemistry, 2019, 58( 5): 312- 329. DOI: 10.1021/acs.biochem.8b00949. [5] BARBIERI I, KOUZARIDES T. Role of RNA modifications in cancer[J]. Nat Rev Cancer, 2020, 20( 6): 303- 322. DOI: 10.1038/s41568-020-0253-2. [6] FURUICHI Y. Discovery of m(7)G-cap in eukaryotic mRNAs[J]. Proc Jpn Acad Ser B Phys Biol Sci, 2015, 91( 8): 394- 409. DOI: 10.2183/pjab.91.394. [7] MALBEC L, ZHANG T, CHEN YS, et al. Dynamic methylome of internal mRNA N7-methylguanosine and its regulatory role in translation[J]. Cell Res, 2019, 29( 11): 927- 941. DOI: 10.1038/s41422-019-0230-z. [8] MARCHAND V, AYADI L, ERNST FGM, et al. AlkAniline-seq: Profiling of m7G and m3C RNA modifications at single nucleotide resolution[J]. Angew Chem Int Ed Engl, 2018, 57( 51): 16785- 16790. DOI: 10.1002/anie.201810946. [9] SONG BW, TANG YJ, CHEN KQ, et al. m7GHub: Deciphering the location, regulation and pathogenesis of internal mRNA N7-methylguanosine(m7G) sites in human[J]. Bioinformatics, 2020, 36( 11): 3528- 3536. DOI: 10.1093/bioinformatics/btaa178. [10] MA JY, HAN H, HUANG Y, et al. METTL1/WDR4-mediated m7G tRNA modifications and m7G codon usage promote mRNA translation and lung cancer progression[J]. Mol Ther, 2021, 29( 12): 3422- 3435. DOI: 10.1016/j.ymthe.2021.08.005. [11] YING XL, LIU BX, YUAN ZS, et al. METTL1-m7 G-EGFR/EFEMP1 axis promotes the bladder cancer development[J]. Clin Transl Med, 2021, 11( 12): e675. DOI: 10.1002/ctm2.675. [12] KATSARA O, SCHNEIDER RJ. m7G tRNA modification reveals new secrets in the translational regulation of cancer development[J]. Mol Cell, 2021, 81( 16): 3243- 3245. DOI: 10.1016/j.molcel.2021.07.030. [13] SUZUKI T. The expanding world of tRNA modifications and their disease relevance[J]. Nat Rev Mol Cell Biol, 2021, 22( 6): 375- 392. DOI: 10.1038/s41580-021-00342-0. [14] ALEXANDROV A, MARTZEN MR, PHIZICKY EM. Two proteins that form a complex are required for 7-methylguanosine modification of yeast tRNA[J]. RNA, 2002, 8( 10): 1253- 1266. DOI: 10.1017/s1355838202024019. [15] BRAUN DA, SHRIL S, SINHA A, et al. Mutations in WDR4 as a new cause of Galloway-Mowat syndrome[J]. Am J Med Genet A, 2018, 176( 11): 2460- 2465. DOI: 10.1002/ajmg.a.40489. [16] SHAHEEN R, ABDEL-SALAM GMH, GUY MP, et al. Mutation in WDR4 impairs tRNA m(7)G46 methylation and causes a distinct form of microcephalic primordial dwarfism[J]. Genome Biol, 2015, 16: 210. DOI: 10.1186/s13059-015-0779-x. [17] TRIMOUILLE A, LASSEAUX E, BARAT P, et al. Further delineation of the phenotype caused by biallelic variants in the WDR4 gene[J]. Clin Genet, 2018, 93( 2): 374- 377. DOI: 10.1111/cge.13074. [18] MICHAUD J, KUDOH J, BERRY A, et al. Isolation and characterization of a human chromosome 21q22.3 gene(WDR4) and its mouse homologue that code for a WD-repeat protein[J]. Genomics, 2000, 68( 1): 71- 79. DOI: 10.1006/geno.2000.6258. [19] BARBIERI I, TZELEPIS K, PANDOLFINI L, et al. Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control[J]. Nature, 2017, 552( 7683): 126- 131. DOI: 10.1038/nature24678. [20] LIU Y, YANG CY, ZHAO Y, et al. Overexpressed methyltransferase-like 1(METTL1) increased chemosensitivity of colon cancer cells to cisplatin by regulating miR-149-3p/S100A4/p53 axis[J]. Aging, 2019, 11( 24): 12328- 12344. DOI: 10.18632/aging.102575. [21] OKAMOTO M, FUJIWARA M, HORI M, et al. tRNA modifying enzymes, NSUN2 and METTL1, determine sensitivity to 5-fluorouracil in HeLa cells[J]. PLoS Genet, 2014, 10( 9): e1004639. DOI: 10.1371/journal.pgen.1004639. [22] PANDOLFINI L, BARBIERI I, BANNISTER AJ, et al. METTL1 promotes let-7 microRNA processing via m7G methylation[J]. Mol Cell, 2019, 74( 6): 1278- 1290.e9. DOI: 10.1016/j.molcel.2019.03.040. [23] ZHANG LS, LIU C, MA HH, et al. Transcriptome-wide mapping of internal N7-methylguanosine methylome in mammalian mRNA[J]. Mol Cell, 2019, 74( 6): 1304- 1316.e8. DOI: 10.1016/j.molcel.2019.03.036. [24] ZHAO YC, KONG LQ, PEI ZQ, et al. m7G methyltransferase METTL1 promotes post-ischemic angiogenesis via promoting VEGFA mRNA translation[J]. Front Cell Dev Biol, 2021, 9: 642080. DOI: 10.3389/fcell.2021.642080. [25] KADUMURI RV, JANGA SC. Epitranscriptomic code and its alterations in human disease[J]. Trends Mol Med, 2018, 24( 10): 886- 903. DOI: 10.1016/j.molmed.2018.07.010. [26] FRAGA MF, URIOL E, BORJA DIEGO L, et al. High-performance capillary electrophoretic method for the quantification of 5-methyl 2′-deoxycytidine in genomic DNA: Application to plant, animal and human cancer tissues[J]. Electrophoresis, 2002, 23( 11): 1677- 1681. DOI: 3.0.CO;2-Z">10.1002/1522-2683(200206)23: 11<1677: AID-ELPS1677>3.0.CO;2-Z. [27] SONG LG, JAMES SR, KAZIM L, et al. Specific method for the determination of genomic DNA methylation by liquid chromatography-electrospray ionization tandem mass spectrometry[J]. Anal Chem, 2005, 77( 2): 504- 510. DOI: 10.1021/ac0489420. [28] ZHENG GQ, QIN YD, CLARK WC, et al. Efficient and quantitative high-throughput tRNA sequencing[J]. Nat Methods, 2015, 12( 9): 835- 837. DOI: 10.1038/nmeth.3478. [29] LI W, LI XY, MA XJ, et al. Mapping the m1A, m5C, m6A and m7G methylation atlas in zebrafish brain under hypoxic conditions by MeRIP-seq[J]. BMC Genomics, 2022, 23( 1): 105. DOI: 10.1186/s12864-022-08350-w. [30] LIN SB, LIU Q, LELYVELD VS, et al. Mettl1/Wdr4-mediated m7G tRNA methylome is required for normal mRNA translation and embryonic stem cell self-renewal and differentiation[J]. Mol Cell, 2018, 71( 2): 244- 255.e5. DOI: 10.1016/j.molcel.2018.06.001. [31] CHIDAMBARANATHAN-REGHUPATY S, FISHER PB, SARKAR D. Hepatocellular carcinoma(HCC): Epidemiology, etiology and molecular classification[J]. Adv Cancer Res, 2021, 149: 1- 61. DOI: 10.1016/bs.acr.2020.10.001. [32] CHEN ZH, ZHU WJ, ZHU SH, et al. METTL1 promotes hepatocarcinogenesis via m7 G tRNA modification-dependent translation control[J]. Clin Transl Med, 2021, 11( 12): e661. DOI: 10.1002/ctm2.661. [33] XIA P, ZHANG H, XU KQ, et al. MYC-targeted WDR4 promotes proliferation, metastasis, and sorafenib resistance by inducing CCNB1 translation in hepatocellular carcinoma[J]. Cell Death Dis, 2021, 12( 7): 691. DOI: 10.1038/s41419-021-03973-5. [34] TIAN QH, ZHANG MF, ZENG JS, et al. METTL1 overexpression is correlated with poor prognosis and promotes hepatocellular carcinoma via PTEN[J]. J Mol Med, 2019, 97( 11): 1535- 1545. DOI: 10.1007/s00109-019-01830-9. [35] ORELLANA EA, LIU Q, YANKOVA E, et al. METTL1-mediated m7G modification of Arg-TCT tRNA drives oncogenic transformation[J]. Mol Cell, 2021, 81( 16): 3323- 3338.e14. DOI: 10.1016/j.molcel.2021.06.031. [36] DAI ZH, LIU HN, LIAO JB, et al. N7-Methylguanosine tRNA modification enhances oncogenic mRNA translation and promotes intrahepatic cholangiocarcinoma progression[J]. Mol Cell, 2021, 81( 16): 3339- 3355.e8. DOI: 10.1016/j.molcel.2021.07.003. [37] HONG ZQ, GOU WX, CUI BQ, et al. Role of the m7G methyltransferase METTL1 in tumours[J]. Chin J Clin Pharmacol Ther, 2023, 28( 1): 93- 100. DOI: 10.12092/j.issn.1009-2501.2023.01.012.洪子强, 苟文曦, 崔百强, 等. m7G甲基转移酶METTL1在肿瘤中的作用[J]. 中国临床药理学与治疗学, 2023, 28( 1): 93- 100. DOI: 10.12092/j.issn.1009-2501.2023.01.012. [38] WANG YT, CHEN J, CHANG CW, et al. Ubiquitination of tumor suppressor PML regulates prometastatic and immunosuppressive tumor microenvironment[J]. J Clin Invest, 2017, 127( 8): 2982- 2997. DOI: 10.1172/JCI89957. [39] HUANG ML, LONG JT, YAO ZJ, et al. METTL1-mediated m7G tRNA modification promotes lenvatinib resistance in hepatocellular carcinoma[J]. Cancer Res, 2023, 83( 1): 89- 102. DOI: 10.1158/0008-5472.CAN-22-0963. -



 PDF下载 ( 888 KB)
PDF下载 ( 888 KB)

 下载:
下载: