地五养肝胶囊对肝癌大鼠模型肝再生微环境影响的表观遗传学机制分析
DOI: 10.12449/JCH260216
Epigenetic mechanism of Diwu Yanggan Capsule in improving liver regeneration microenvironment in a rat model of liver cancer
-
摘要:
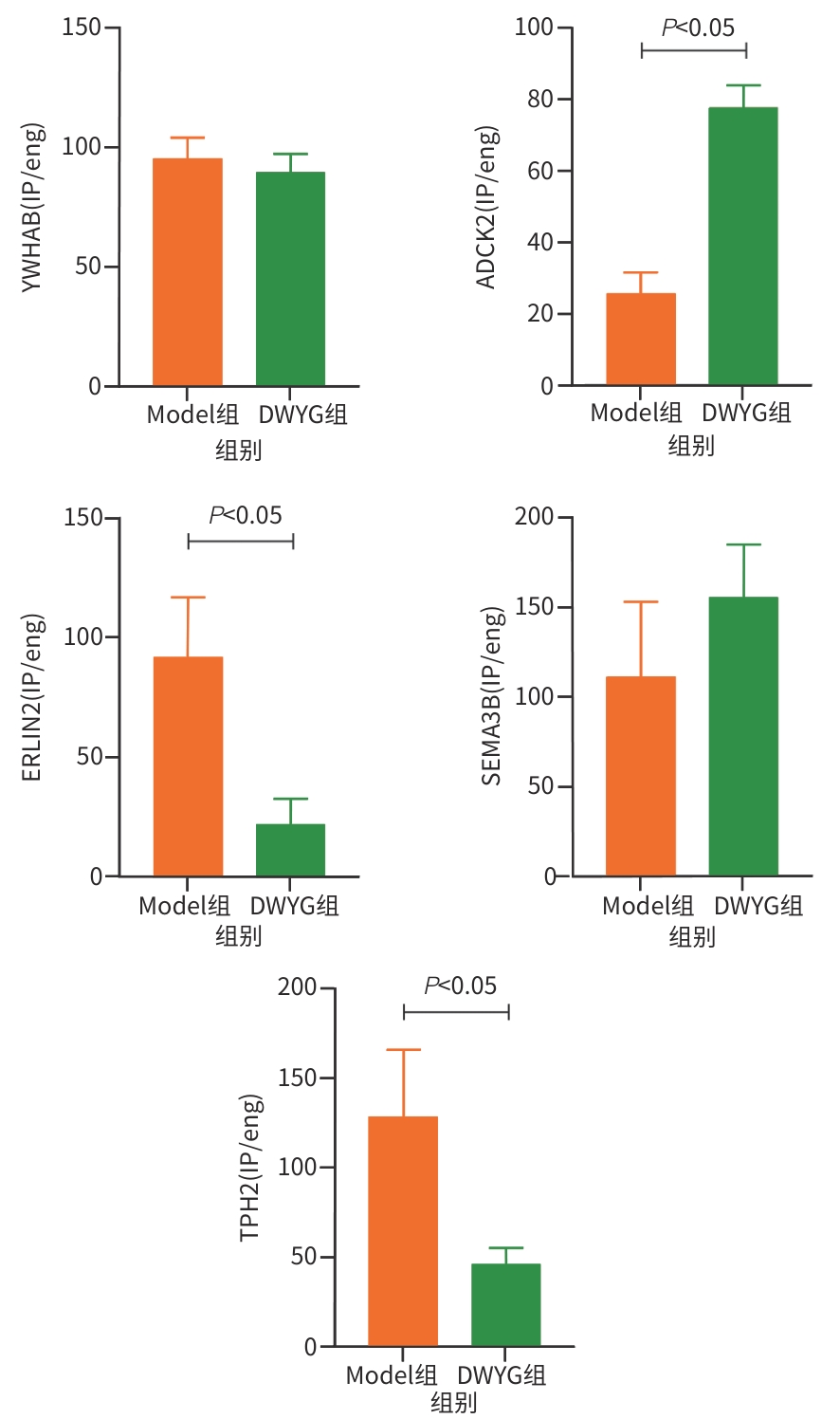
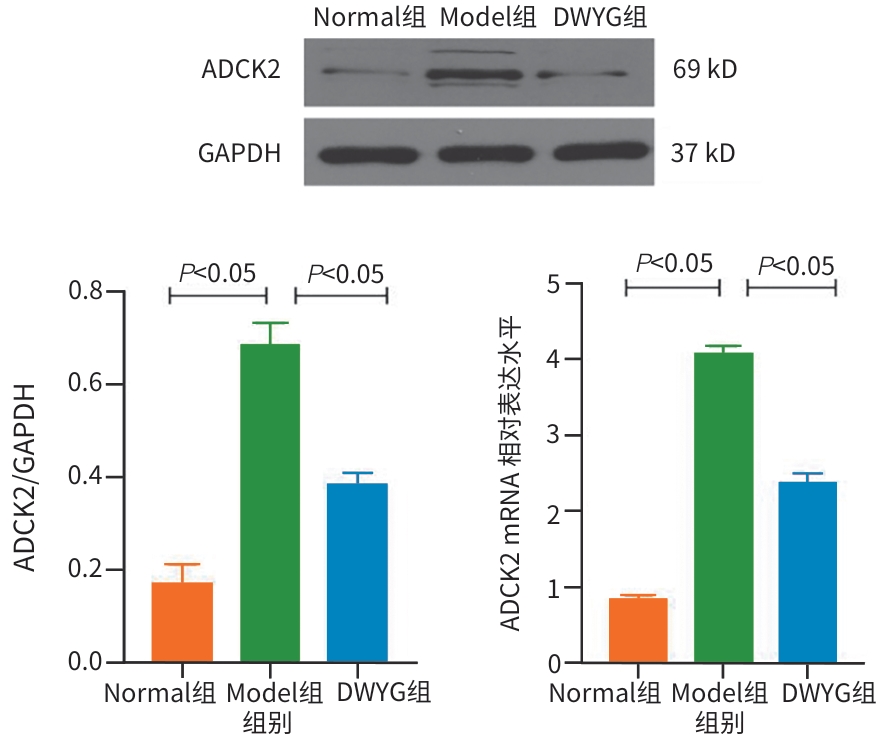
目的 探讨地五养肝胶囊通过调控DNA甲基化,改善肝癌大鼠模型肝再生微环境的表观遗传学机制,为临床提供科学用药依据。 方法 选取48只无特定病原体级SD大鼠,按照随机数字表法分为正常组、模型组和地五养肝胶囊组,每组16只。采用Solt-Farber二步法复制肝癌大鼠模型,地五养肝胶囊组大鼠给予地五养肝胶囊750 mg·kg⁻¹·d⁻¹灌胃处理,正常组及模型组给予等量生理盐水灌胃处理,连续干预16周后采集各组大鼠肝组织,使用DNA甲基化芯片分析各组大鼠肝组织DNA甲基化变化情况,并运用基因本体论分析(GO分析)及京都基因与基因组百科全书分析(KEGG分析)对数据进行整合分析。进一步运用甲基化DNA免疫共沉淀-聚合酶链式反应(MeDIP-PCR)技术检测各组大鼠肝组织中YWHAB、ADCK2、ERLIN2、SEMA3B和TPH2等候选差异甲基化基因变化情况,并运用Western Blot及RT-qPCR验证关键甲基化基因表达情况。计量资料两组间比较采用成组t检验,多组间比较采用单因素方差分析,进一步两两比较采用LSD-t检验。 结果 DNA甲基化芯片分析发现,与正常组相比,肝癌模型组大鼠肝组织中共检测到2 422个基因启动子区域发生显著甲基化改变。进一步通过GO功能富集与KEGG通路分析显示,这些差异甲基化基因显著富集于类固醇激素生物合成、药物代谢-细胞色素P450等代谢相关通路。与模型组相比,地五养肝胶囊组肝组织中检测到1 650个基因启动子甲基化状态发生显著逆转,其中KEGG富集分析提示,这些基因主要参与钙离子信号通路、cAMP信号通路及细胞外因子信号通路等与细胞增殖、凋亡及微环境调节密切相关的途径。与模型组比较,地五养肝胶囊干预组中ADCK2基因的启动子甲基化水平显著升高(P<0.05),ERLIN2及TPH2基因的启动子甲基化水平显著降低(P值均<0.05)。与模型组相比,地五养肝胶囊干预组中ADCK2 mRNA与蛋白表达显著降低(P值均<0.05)。 结论 肝组织DNA甲基化异常参与肝癌的发生与发展,地五养肝胶囊影响DNA甲基化水平是其防治肝癌的重要表观遗传学机制。 Abstract:Objective To investigate the epigenetic mechanism of Diwu Yanggan Capsule in improving liver regeneration microenvironment in a rat model of liver cancer by regulating DNA methylation, and to provide a basis for scientific clinical medication. Methods A total of 48 specific pathogen-free Sprague-Dawley rats were divided into normal group, model group, and Diwu Yanggan Capsule group using a random number table, with 16 rats in each group. The Solt-Farber two-step method was used to establish a rat model of liver cancer. The rats in the Diwu Yanggan Capsule group were given Diwu Yanggan Capsule at a dose of 750 mg/kg/d by gavage, and those in the normal group and the model group were given an equal volume of normal saline by gavage. Liver tissue samples were collected from each group of rats after 16 weeks of continuous intervention; DNA methylation chips were used to analyze the change in DNA methylation in liver tissue, and gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were used for data analysis. In addition, the MeDIP-PCR technique was used to detect the changes in candidate differentially methylated genes such as YWHAB, ADCK2, ERLIN2, SEMA3B, and TPH2 in the liver tissue of rats, and Western blot and RT-qPCR were used to verify the expression of key methylated genes. The independent-samples t test was used for comparison of continuous data between two groups, and a one-way analysis of variance was used for comparison between multiple groups, while the least significant difference t-test was used for further comparison between two groups. Results The DNA methylation chip analysis showed that compared with the normal group, the model group had significant methylation changes in the promoter region of 2 422 genes in liver tissue of rats. The GO functional enrichment analysis and the KEGG pathway enrichment analysis showed that these differentially methylated genes were significantly enriched in metabolic pathways such as steroid hormone biosynthesis and drug metabolism-cytochrome P450. Compared with the model group, the Diwu Yanggan Capsule group had significant reversal of promoter methylation in 1 650 genes, and the KEGG enrichment analysis showed that these genes were mainly involved in the pathways closely associated with cell proliferation, apoptosis, and microenvironment regulation, such as the calcium ion signaling pathway, the cAMP signaling pathway, and the extracellular factor signaling pathway. Compared with the model group, the Diwu Yanggan Capsule group had a significant increase in the promoter methylation level of the ADCK2 gene (P<0.05) and significant reductions in the promoter methylation levels of the ERLIN2 and TPH2 genes (all P<0.05). Compared with the model group, the Diwu Yanggan Capsule group had significant reductions in the mRNA expression levels and the protein expression levels of the ADCK2 (all P<0.05). Conclusion Abnormal DNA methylation in liver tissue participates in the development and progression of liver cancer. The effect of Diwu Yanggan Capsule on DNA methylation level is an important epigenetic mechanism for its effect in the prevention and treatment of liver cancer. -
注: a,各组DNA样本OD260/OD230比值分布柱状图;b:各组DNA样本OD260/OD280比值分布柱状图;c,组内样本DNA数据相关性散点图,展示同组(如Normal组)不同重复样本间的DNA数据重复性;d,组间样本DNA数据相关性散点图,展示不同组(如Normal组vs Model组)样本间的DNA数据关联度;e,样本间DNA数据相关系数矩阵图,图中数值为样本间的Pearson相关系数(越接近1,代表样本数据的一致性、重复性越高)。
图 1 各组大鼠肝癌组织提取DNA质量控制数据及各组间相关矩阵分析分布图
Figure 1. Quality control data of DNA extracted from liver cancer tissues of each group and distribution map of correlation analysis among different groups
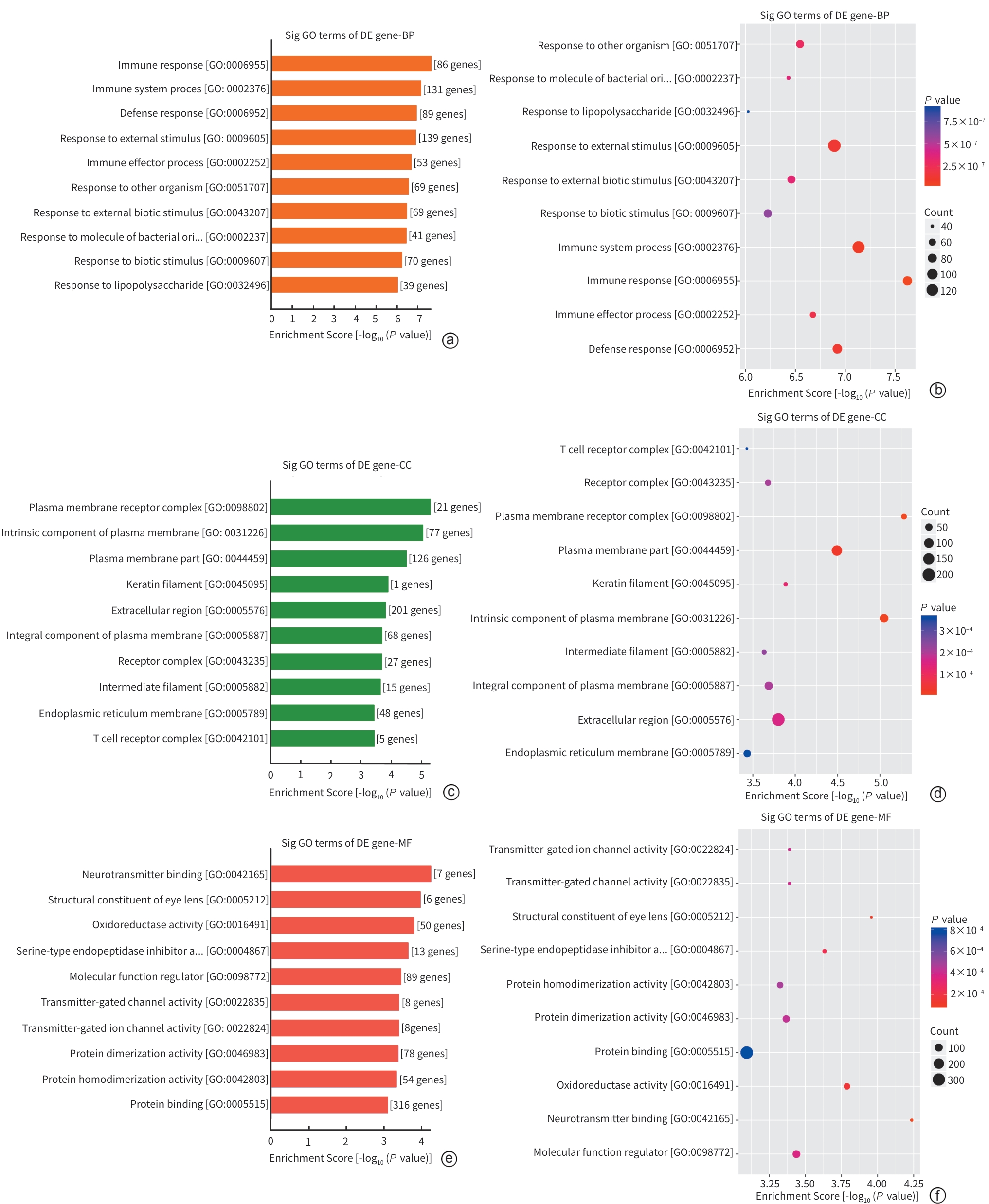
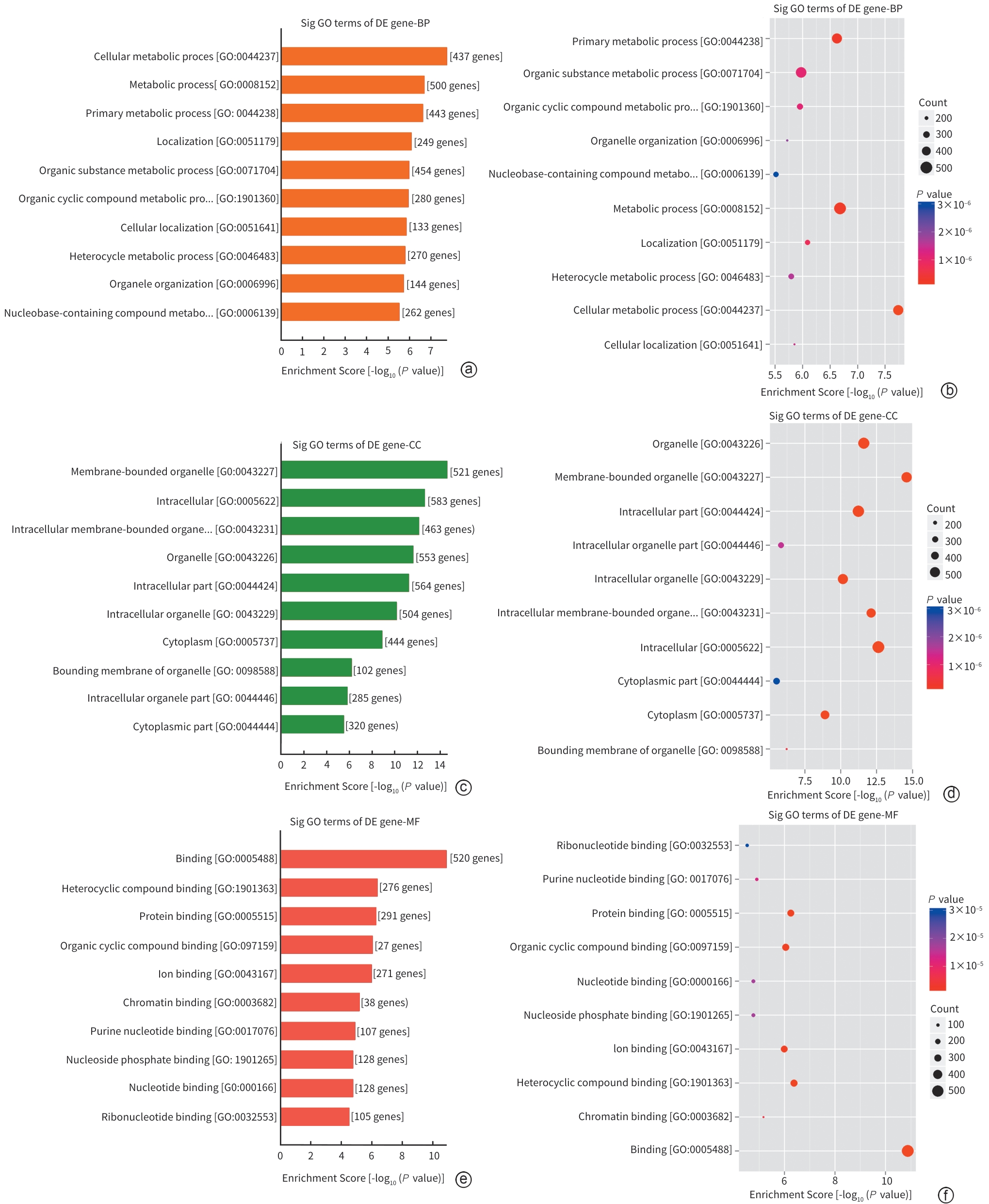
注: a,差异基因的生物过程显著富集GO term柱状图;b,差异基因的生物过程显著富集GO term气泡图;c,差异基因的细胞组分显著富集GO term柱状图;d,差异基因的细胞组分显著富集GO term气泡图;e,差异基因的分子功能显著富集GO term柱状图;f,差异基因的分子功能显著富集GO term气泡图。
图 4 DWYG组与Model组差异启动子甲基化基因GO分析
Figure 4. GO analysis of differentially methylated promoter genes between the Diwu Yanggan Capsule group and the model group
表 1 引物序列
Table 1. Primer sequence
基因名 引物序列(5'-3') 退火温
度(℃)产物长
度(bp)YWHAB 60 238 F 5'-CCGCCCTCTCTTCTTCCTC-3' R 5'-CGCTCTAGCCCAAGTCCC-3' ADCK2 60 142 F 5'-AGGCTTTTGCCCTGGTTC-3' R 5'-GCACGGAGCCTACAGTCAG-3' ERLIN2 60 146 F 5'-ACCCCACGCCACCTCTTG-3' R 5'-TCACTCAGCTCGCCCACC-3' SEMA3B 60 173 F 5'-CACCCTCCTCTGTAACCTCC-3' R 5'-TGCCCTCTGAAATGACCAC-3' TPH2 60 164 F 5'-AGGCAGACAGAAAGGAGGGA-3' R 5'-CTTGGCGGCTTAACAGTGAG-3' -
[1] BROWN ZJ, TSILIMIGRAS DI, RUFF SM, et al. Management of hepatocellular carcinoma: A review[J]. JAMA Surg, 2023, 158( 4): 410- 420. DOI: 10.1001/jamasurg.2022.7989. [2] SUNG H, FERLAY J, SIEGEL RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71( 3): 209- 249. DOI: 10.3322/caac.21660. [3] ZHOU J, SUN HC, WANG Z, et al. Guidelines for the diagnosis and treatment of primary liver cancer(2022 edition)[J]. Liver Cancer, 2023, 12( 5): 405- 444. DOI: 10.1159/000530495. [4] RICH NE, YOPP AC, SINGAL AG. Medical management of hepatocellular carcinoma[J]. J Oncol Pract, 2017, 13( 6): 356- 364. DOI: 10.1200/JOP.2017.022996. [5] CHEN WQ. Cancer statistics: Updated cancer burden in China[J]. Chin J Cancer Res, 2015, 27( 1): 1. DOI: 10.3978/j.issn.1000-9604.2015.02.07. [6] TAHMASEBI BIRGANI M, CARLONI V. Tumor microenvironment, a paradigm in hepatocellular carcinoma progression and therapy[J]. Int J Mol Sci, 2017, 18( 2): 405. DOI: 10.3390/ijms18020405. [7] SAS Z, CENDROWICZ E, WEINHÄUSER I, et al. Tumor microenvironment of hepatocellular carcinoma: Challenges and opportunities for new treatment options[J]. Int J Mol Sci, 2022, 23( 7): 3778. DOI: 10.3390/ijms23073778. [8] CLEVERS H. The cancer stem cell: Premises, promises and challenges[J]. Nat Med, 2011, 17( 3): 313- 319. DOI: 10.1038/nm.2304. [9] WU K, DING J, CHEN C, et al. Hepatic transforming growth factor beta gives rise to tumor-initiating cells and promotes liver cancer development[J]. Hepatology, 2012, 56( 6): 2255- 2267. DOI: 10.1002/hep.26007. [10] LI HM, ZHANG LS. Liver regeneration microenvironment of hepatocellular carcinoma for prevention and therapy[J]. Oncotarget, 2017, 8( 1): 1805- 1813. DOI: 10.18632/oncotarget.12101. [11] LI HM. Microcirculation of liver cancer, microenvironment of liver regeneration, and the strategy of Chinese medicine[J]. Chin J Integr Med, 2016, 22( 3): 163- 167. DOI: 10.1007/s11655-016-2460-y. [12] LI HM, YE ZH. Microenvironment of liver regeneration in liver cancer[J]. Chin J Integr Med, 2017, 23( 7): 555- 560. DOI: 10.1007/s11655-017-2806-0. [13] WANG MG, YE QL, MAO DW, et al. Research progress in liver-regenerating microenvironment and DNA methylation in hepatocellular carcinoma: The role of traditional Chinese medicine[J]. Med Sci Monit, 2020, 26: e920310. DOI: 10.12659/MSM.920310. [14] LI Y, LI HM, LI ZC, et al. Ingredients, anti-liver cancer effects and the possible mechanism of DWYG formula based on network prediction[J]. Onco Targets Ther, 2020, 13: 4213- 4227. DOI: 10.2147/OTT.S238901. [15] LI HM, YE ZH, GAO X, et al. Diwu Yanggan capsule improving liver histological response for patients with HBeAg-negative chronic hepatitis B: A randomized controlled clinical trial[J]. Am J Transl Res, 2018, 10( 5): 1511- 1521. [16] ZHAO BB, YE ZH, GAO X, et al. Diwu Yanggan modulates the Wnt/β-catenin pathway and inhibits liver carcinogenesis signaling in 2-AAF/PH model rats[J]. Curr Med Sci, 2019, 39( 6): 913- 919. DOI: 10.1007/s11596-019-2123-2. [17] YE ZH, GAO X, ZHAO BB, et al. Diwu Yanggan capsule inhibits the occurrence and development of liver cancer in the Solt-Farber rat model by regulating the Ras/Raf/Mek/Erk signaling pathway[J]. Am J Transl Res, 2018, 10( 11): 3797- 3805. [18] ZHAO BB, LI HM, GAO X, et al. The herbal compound“Diwu Yanggan” modulates liver regeneration by affecting the hepatic stem cell microenvironment in 2-acetylaminofluorene/partial hepatectomy rats[J]. Evid Based Complement Alternat Med, 2015, 2015: 468303. DOI: 10.1155/2015/468303. [19] VIERTHALER M, SUN Q, WANG YM, et al. ADCK2 knockdown affects the migration of melanoma cells via MYL6[J]. Cancers, 2022, 14( 4): 1071. DOI: 10.3390/cancers14041071. [20] VÁZQUEZ-FONSECA L, SCHAEFER J, NAVAS-ENAMORADO I, et al. ADCK2 haploinsufficiency reduces mitochondrial lipid oxidation and causes myopathy associated with CoQ deficiency[J]. J Clin Med, 2019, 8( 9): 1374. DOI: 10.3390/jcm8091374. [21] ZHANG JZ, LIU J, XU YX, et al. Identification of the mitochondrial protein ADCK2 as a therapeutic oncotarget of NSCLC[J]. Int J Biol Sci, 2022, 18( 16): 6163- 6175. DOI: 10.7150/ijbs.78354. [22] HERNÁNDEZ-CAMACHO JD, FERNÁNDEZ-AYALA DJM, VICENTE-GARCÍA C, et al. Calorie restriction rescues mitochondrial dysfunction in Adck2-deficient skeletal muscle[J]. Front Physiol, 2022, 13: 898792. DOI: 10.3389/fphys.2022.898792. [23] XU Y, WANG L, ZHANG H, et al. ADCK2 regulates protein phosphorylation and membrane composition in mammalian cells[J]. Biochem Biophys Res Commun, 2020, 529( 4): 987- 993. DOI: 10.1016/j.bbrc.2020.07.054. [24] SCHOOLMEESTERS A, BROWN DD, FEDOROV Y. Kinome-wide functional genomics screen reveals a novel mechanism of TNFα-induced nuclear accumulation of the HIF-1α transcription factor in cancer cells[J]. PLoS One, 2012, 7( 2): e31270. DOI: 10.1371/journal.pone.0031270. -



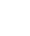 PDF下载 ( 9880 KB)
PDF下载 ( 9880 KB)
 下载:
下载: